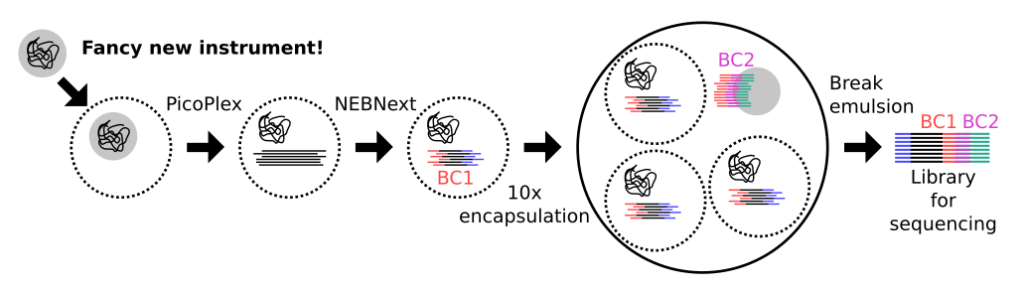
A fundamental question in microbiome research involves determining which organisms are part of a given microbial community. Despite major breakthroughs in shotgun metagenomic sequencing (SMS) and bioinformatic analysis methods, determining “who is there” in a microbial community remains extremely challenging: de novo assembly of sequencing reads generated via SMS typically results in a collection of contigs that cannot be grouped into strain-level genomes. Here, the postdoc will develop a high-throughput method, which can reconstruct whole microbial genomes from microbiome samples at the single-cell level. Following prototyping, the method will be used to characterize clinical oral microbiome samples collected from a cohort of Swedish patients, with the goal of understanding why some humans develop chronic oral infections, while others do not.
This postdoc will be housed in IceLab and hosted by the Departments of Clinical Microbiology, Molecular Biology, and Odontology and supervised by a multidisciplinary team with complementing expertise in microbiology, single-cell technology development, bioinformatics, computer science, and dentistry.
Specific Qualifications
To qualify for the fellowship, the candidate should have a PhD degree (or a foreign degree that is deemed equivalent), preferably in (micro)biology, molecular biology, (bio)chemistry, bioinformatics, engineering, physics, computer science, or a related field; candidates from other fields with high-throughput (meta)genomic and/or single-cell sequencing experience (e.g., sequencing library preparation, analysis of Illumina or long-read sequencing data; see below) are also strongly encouraged to apply.
The project includes both generation and analysis of single-cell metagenomic data; as such, we are ideally looking for someone with “moist lab” experience (i.e., a mix of wet lab and computational skills). However, we strongly welcome applications from highly skilled, motivated wet or dry lab scientists who are willing to collaborate with our research groups to learn new skills.
The ideal candidate has:
- Proficiency in written and spoken English.
- A desire to work in a diverse, inclusive, interdisciplinary, and highly collaborative environment.
We welcome applications from candidates with combinations of any of the following skills:
- Experience working with DNA or RNA in any form, especially within the context of advanced cloning methods and/or preparing DNA/RNA for high-throughput sequencing.
- Experience working with bacterial or eukaryotic cells and associated methods (e.g., flow cytometry).
- Experience developing or optimizing protocols for high-throughput sequencing.
- Experience using single-cell instruments and/or robotics for wet lab automation.
- Experience analyzing Illumina or long-read sequencing data, preferably single-cell data derived from any organism or (meta)genomic/(meta)transcriptomic data derived from microbes.
- Proficiency in Bash shell scripting and a relevant programming language (e.g., Python, R, C, or Java).
- Knowledge of statistics and machine learning methods.
- Knowledge of data structures and algorithms.
Contact Information
Dr. Laura Carroll, laura.carroll@umu.se
Dr. Johan Henriksson, johan.henriksson@umu.se
The IceLab Multidisciplinary Postdoctoral Program
The under-explored terrain between traditional disciplines is full of fascinating and impactful research questions. At IceLab, we promote and facilitate transdisciplinary collaborations – with a focus on cutting-edge research that integrates theoretical, computational, and empirical work.
We will welcome you to IceLab with genuine support by creative researchers working on a multitude of interdisciplinary problems. You will participate in both professionally and personally rewarding and entertaining activities aimed at training a new kind of researcher. A multidisciplinary team of researchers with complementary expertise will supervise each postdoc.
The two-year postdoc fellowships are financed by the Kempe foundations and are part of the IceLab Multidisciplinary Postdoctoral Program. A fellowship amounts to 2 years funding: 28,000 SEK per month. The scholarships is tax-free. Application deadline September 21, 2023. Start winter/spring 2024 (exact start date according to agreement).
Formal qualifications
To qualify as a postdoctoral scholarship holder, the postdoctoral fellow is required to have completed a doctoral degree or a foreign degree deemed equivalent to a doctoral degree. This qualification requirements must be fulfilled no later than at the time of the decision about scholarship recipient.
Priority should be given to candidates who completed their doctoral degree, according to what is stipulated in the paragraph above, no later than three years prior. If there are special reasons, candidates who completed their doctoral degree prior to that may also be eligible. Special reasons include absence due to illness, parental leave, appointments of trust in trade union organizations, military service, or similar circumstances, as well as clinical practice or other forms of appointment/assignment relevant to the subject area.
Candidates should have experience in computational and quantitative modeling or an interest in learning these skills. Personal qualities such as collaboration, communication, strong drive and motivation, critical thinking abilities, creativity and analytical skills are essential. You should be able to take on the research independently and as part of a team. Good knowledge of oral and written English is required.
Application
A full application should include:
- A cover letter clearly stating which project or projects you are particularly interested in and summarizing your qualifications, your scientific interests, and your motives for applying (max 2 pages),
- A curriculum vitae (CV) with publication list,
- Certified copy of doctoral degree certificate,
- Certified copies of other diplomas, list of completed academic courses and grades,
- Copy of doctoral thesis,
- Copies of relevant publications,
- Contact information for at least two reference persons,
- Other documents that the applicant wishes to claim.
To apply, follow this link for further instructions.
We look forward to receiving your application.
